Unified analytics engine for large-scale data processing (mostly in-memory).
- Main Features
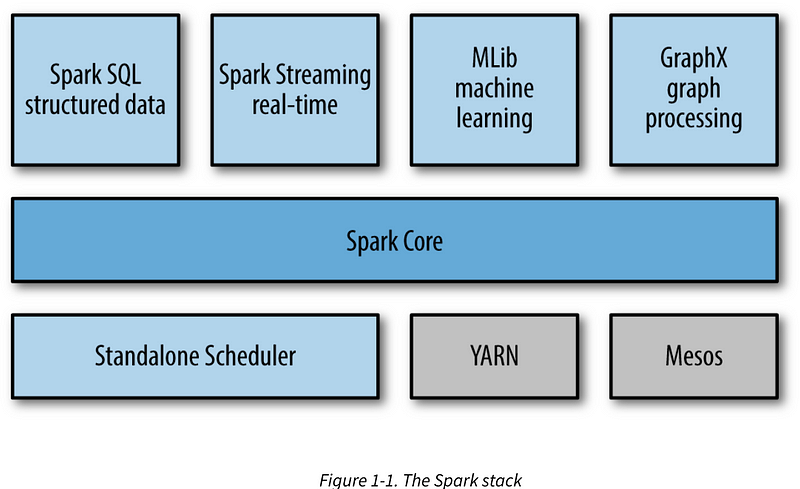
- Components
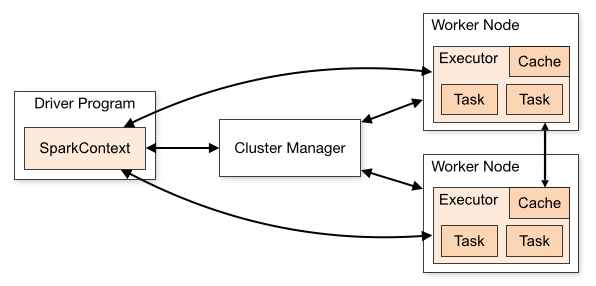
- Architecture
- RDD
- RDD Operations
- How To Start Shell
- How to Run Application
- Managing Partitions
- RDD Persistence
- RDD Checkpoint
- Understanding Closures
- Few Don'ts
- Working with Key-Value Pairs
- Shared Variables
- Spark SQL
- Dataset
- Internals
- Performance Tuning
- Best Practices
- Partition and Executor Parameter Estimation
- Extras
- TODO
- References
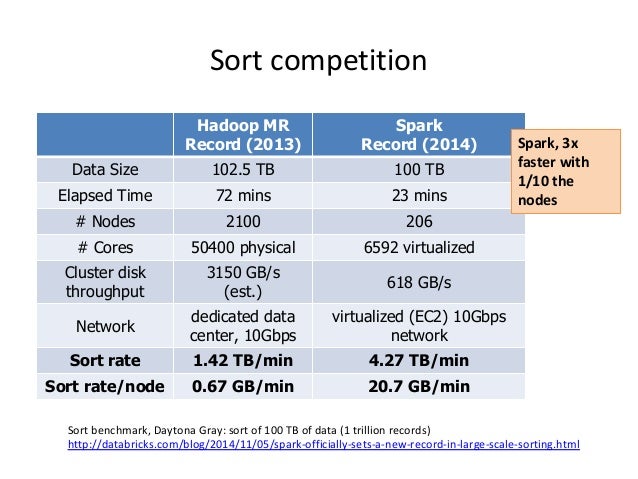
- Faster than Map-Reduce
- Runs on Hadoop, Apache Mesos, Kubernetes, standalone, or in the cloud. It can access diverse data sources
- Write applications quickly in Java, Scala, Python, R, and SQL
It's actually easier to write code in spark
Dataset<Row> df = session.read().json("logs.json");
df.where("age > 21").select("name.first").show();
True. Isn't it? 😎
Spark current version - 2.4.2 (as of April'19)
Platform versions
| Language | Version |
|---|---|
| Java | 8+ |
| Python | c2.7+/3.1+ |
| R | 3.1+ |
| Scala | 2.11.x |
Spark is packaged with a built-in cluster manager called the Standalone cluster manager. Spark also works with Hadoop YARN and Apache Mesos.
A driver is the process where the main() method of your program runs. It is the process running the user code that creates a SparkContext, creates RDDs and performs transformations and actions.
A driver will do two major tasks:
Converting a user program into tasks
Spark program implicitly creates a logical DAG of operations. When the driver runs, it converts this logical graph into a physical execution plan. Spark performs several optimizations, such as “pipelining” map transformations together to merge them, and converts the execution graph into a set of stages. Each stage, in turn, consists of multiple tasks.
Scheduling tasks on executors
The Spark driver will look at the current set of executors and try to schedule each task in an appropriate location, based on data placement.
Spark executors are worker processes responsible for running the individual tasks in a given Spark job. Executors provide in-memory storage for RDDs that are cached by user programs, through a service called the Block Manager that lives within each executor
Resilient distributed dataset (RDD), which is a collection of elements partitioned across the nodes of the cluster that can be operated on in parallel. RDDs automatically recover from node failures. It can be created parallelizing a collection
List<Integer> data = Arrays.asList(1, 2, 3, 4, 5, 6, 7, 8, 9, 10);
JavaRDD<Integer> distData = sc.parallelize(data, 2);
Or reading a file
String path = "/home/devender/data/sample.txt";
sc.textFile(path,2);
- Transformation
- Action
All transformations in Spark are lazy and are only computed when an action requires a result to be returned to the driver program.
Different transformations:
- map(func)
- filter(func)
- union(otherDataset)
- intersection(otherDataset)
- coalesce(numPartitions)
- repartition(numPartitions)
- distinct([numPartitions]))
and many more. For details check spark transformations.
By default, each transformed RDD may be recomputed each time you run an action on it. However, you may also persist an RDD in memory using the persist (or cache) method.
Different actions:
- Collect()
- count()
- first()
- take(n)
and many more. For details check spark actions.
Sample Program
import java.util.Arrays;
import java.util.List;
import org.apache.spark.SparkConf;
import org.apache.spark.api.java.JavaRDD;
import org.apache.spark.api.java.JavaSparkContext;
public class TestRdd {
public static void main(String[] args) {
SparkConf conf = new SparkConf().setAppName("sample-app").setMaster("local[2]");
try (JavaSparkContext sc = new JavaSparkContext(conf);) {
List<Integer> data = Arrays.asList(1, 2, 3, 4, 5);
JavaRDD<Integer> rdd = sc.parallelize(data);
JavaRDD<Integer> squareRdd = rdd.map(s -> s * s);
Integer sum = squareRdd.reduce((a, b) -> a + b);
System.out.println("sum of square of all the elements - " + sum);
}
}
}
Run Spark interactively (scala shell)
./bin/spark-shell --master local[2]
in python interpretor
./bin/pyspark --master local[2]
in R interpretor
./bin/sparkR --master local[2]
bin/spark-submit --master spark://host:7077 --executor-memory 10g spark-app.jar
Different values for master:
- Standalone mode (
spark://host:port) - Mesos (
mesos://host:port) - yarn (
yarn) - local (
local,local[N],local[*])
Common flags for spark-submit:
--master--deploy-mode--class--name--jars--files--py-files--executor-memory--driver-memory
Example:
./bin/spark-submit \
--master yarn \
--deploy-mode cluster \
--name "Example Program" \
--num-executors 40 \
--executor-memory 10g \
--class com.example.MyApp
my-project.jar "options" "to your application" "go here"
At a high level, every Spark application consists of a driver program that runs the user’s main function and executes various parallel operations on a cluster.
By default, Spark creates one partition for each block of the file (blocks being 128MB by default in HDFS), but you can also ask for a higher number of partitions by passing a larger value.
Spark will run one task for each partition of the cluster. Typically you want 2-4 partitions for each CPU in your cluster. Normally, Spark tries to set the number of partitions automatically based on your cluster. The number of partitions can be configured while creating and transforming RDD.
Hash Partitioning in Spark
Hash Partitioning attempts to spread the data evenly across various partitions based on the key. Object.hashCode method is used to determine the partition in Spark as partition = key.hashCode () % numPartitions.
Range Partitioning in Spark
Some Spark RDDs have keys that follow a particular ordering, for such RDDs, range partitioning is an efficient partitioning technique. In range partitioning method, tuples having keys within the same range will appear on the same machine. Keys in a range partitioner are partitioned based on the set of sorted range of keys and ordering of keys.
When you persist an RDD, each node stores any partitions of it that it computes in memory and reuses them in other actions on that dataset (or datasets derived from it). This allows future actions to be much faster (often by more than 10x). Caching is a key tool for iterative algorithms and fast interactive use.
You can mark an RDD to be persisted using the persist() or cache() methods on it.
Storage Level -
- MEMORY_ONLY
- MEMORY_AND_DISK
- MEMORY_ONLY_SER
- MEMORY_AND_DISK_SER
- DISK_ONLY
- MEMORY_ONLY_2
- MEMORY_AND_DISK_2
- OFF_HEAP
Spark automatically monitors cache usage on each node and drops out old data partitions in a least-recently-used (LRU) fashion. If you would like to manually remove an RDD instead of waiting for it to fall out of the cache, use the RDD.unpersist() method.
Checkpointing is a process of truncating RDD lineage graph and saving it to a reliable distributed (HDFS) or local file system. Checkpointing can be done using RDD.checkpoint() method.
Checkpoint directory should be set before checkpointing.
SparkContext.setCheckpointDir(directory)
Checkpointing can be used to truncate the logical plan of this Dataset, which is especially useful in iterative algorithms where the plan may grow exponentially.
By default, checkpointing is done eagerly. It can be controlled using checkpoint(boolean eager) method.
Prior to execution, Spark computes the task’s closure. The closure is those variables and methods which must be visible for the executor to perform its computations on the RDD. More details here.
int counter = 0;
JavaRDD<Integer> rdd = sc.parallelize(data);
// Wrong: Don't do this!!
rdd.foreach(x -> counter += x);
println("Counter value: " + counter);
To ensure well-defined behavior in these sorts of scenarios one should use an Accumulator.
-
Don't print like that:
rdd.foreach(println)It will print on executor, not drivers.
-
Don't collection very large RDD
rdd.collect().foreach(println)It will bring complete collection in memory. This can cause the driver to run out of memory. A safer approach is to use the
take(). For example,rdd.take(100).foreach(println).
Few spark operations are available on key-value pairs. The most common ones are distributed shuffle operations, such as groupByKey, reduceByKey, aggregateByKey.
To count how many times each line of text occurs in a file:
Sample Data:
words.txt
apple
banana
grapes
apple
orange
pear
orange
grapes
Spark Program
JavaRDD<String> lines = sc.textFile("file:///home/devender/data/words.txt");
JavaPairRDD<String, Integer> pairRdd = lines.mapToPair(s -> new Tuple2(s, 1));
pairRdd.collect().forEach(System.out::println);
JavaPairRDD<String, Iterable<Integer>> groupedPairRdd = pairRdd.groupByKey();
groupedPairRdd.collect().forEach(System.out::println);
JavaPairRDD<String, Integer> reducedPairRdd = pairRdd.reduceByKey((a, b) -> a + b);
reducedPairRdd.collect().forEach(System.out::println);
Output:
PairRdd:
(apple,1)
(banana,1)
(grapes,1)
(apple,1)
(orange,1)
(pear,1)
(orange,1)
(grapes,1)
Group by key output:
(pear,[1])
(orange,[1, 1])
(apple,[1, 1])
(grapes,[1, 1])
(banana,[1])
Reduce by key output:
(pear,1)
(orange,2)
(apple,2)
(grapes,2)
(banana,1)
Note: when using custom objects as the key in key-value pair operations, you must be sure that a custom equals() method is accompanied with a matching hashCode() method.
Another abstraction in Spark is shared variables that can be used in parallel operations. By default, when Spark runs a function in parallel as a set of tasks on different nodes, it ships a copy of each variable used in the function to each task. Sometimes, a variable needs to be shared across tasks, or between tasks and the driver program. Spark supports two types of shared variables: broadcast variables, which can be used to cache a value in memory on all nodes, and accumulators, which are variables that are only "added" to, such as counters and sums.
There are two types of shared variables:
- broadcast variables
- accumulators
Broadcast variables are immutable shared variables which are cached on each worker nodes on a Spark cluster. Broadcast variables allow the programmer to keep a read-only variable cached on each machine rather than shipping a copy of it with tasks.
Broadcast<int[]> broadcastVar = sc.broadcast(new int[] {1, 2, 3});
broadcastVar.value();
// returns [1, 2, 3]
If you have a huge array that is accessed from Spark Closures, for example, some reference data, this array will be shipped to each spark node with closure.
Why use Broadcast Variables?
For example, if you have 10 nodes cluster with 100 partitions (10 partitions per node), this Array will be distributed at least 100 times (10 times to each node). If you use broadcast it will be distributed once per node using efficient p2p protocol 😎
Accumulators are variables that are only "added" to through an associative and commutative operation and can therefore be efficiently supported in parallel. Tasks running on a cluster can then add to it using the add method. Executors can't read the accumulator’s value. Only the driver program can read the accumulator’s value, using its value method.
LongAccumulator accum = jsc.sc().longAccumulator();
sc.parallelize(Arrays.asList(1, 2, 3, 4)).foreach(x -> accum.add(x));
accum.value();
// returns 10
Now let's fix the example from understanding-closures
List<Integer> data = Arrays.asList(1, 2, 3, 4);
JavaRDD<Integer> rdd = sc.parallelize(data);
LongAccumulator accum = sc.sc().longAccumulator();
rdd.foreach(i -> accum.add(i));
System.out.println("Counter value: " + accum.value());
Spark SQL is a Spark module for structured data processing. Interaction with Spark SQL can be done via SQL and Dataset API. Internally, the same execution engine is used. So, developers can easily switch between different APIs 😍
Also, check spark sql built-in functions.
Dataset is a distributed collection of data that provides the benefits of RDDs with the benefits of Spark SQL’s optimized execution engine. Dataset can be constructed from JVM objects and then manipulated using functional transformations.
Check Javadoc for various transformations and actions on Dataset.
Spark 2.0 provides built-in support for Hive features including the ability to write queries using HiveQL, access to Hive UDFs, and the ability to read data from Hive tables
A DataFrame is a Dataset organized into named columns and is represented by Dataset.
Create SparkSession
import org.apache.spark.sql.SparkSession;
SparkSession spark = SparkSession
.builder()
.appName("Java Spark SQL basic example")
.config("spark.some.config.option", "some-value")
.getOrCreate();
Creating Dataframe
import org.apache.spark.sql.Dataset;
import org.apache.spark.sql.Row;
Dataset<Row> df = spark.read().json("examples/src/main/resources/people.json");
// Displays the content of the DataFrame to stdout
df.show();
Creating Dataset
Instead of using Java serialization or Kryo Datasets use a specialized Encoder to serialize the objects for processing or transmitting over the network.
public static class Person implements Serializable {
private String name;
private int age;
// getters and setters
}
// Create an instance of a Bean class
Person person = new Person();
person.setName("Andy");
person.setAge(32);
// Encoders are created for Java beans
Encoder<Person> personEncoder = Encoders.bean(Person.class);
Dataset<Person> javaBeanDS = spark.createDataset(
Collections.singletonList(person),
personEncoder
);
javaBeanDS.show();
Create from a Dataframe
Dataframe.as(Encoder) can be used.
String path = "examples/src/main/resources/people.json";
Dataset<Person> peopleDS = spark.read().json(path).as(personEncoder);
peopleDS.show();
Apart from that, check commonly used fuctions for Dataframe. These are very helpful.
Untyped Dataset (Dataframe) Operations
df.select("name").show();
df.select(col("name"), col("age").plus(1)).show();
df.filter(col("age").gt(21)).show();
Run SQL Queries
df.createOrReplaceTempView("people");
Dataset<Row> sqlDF = spark.sql("SELECT * FROM people");
sqlDF.show();
Run SQL directly on files
Dataset<Row> sqlDF =
spark.sql("SELECT * FROM parquet.`examples/src/main/resources/users.parquet`");
spark.read().json(path)
spark.read().csv(path)
spark.read().jdbc(url, table, properties)
Remembering sqoop? Me too 😊
By default, spark will read data from JDBC and write in one partition. But we can specify the partition column, lowerBound and upperBound to perform the task in parallel ![]()
spark.read().jdbc(url, table, columnName, lowerBound, upperBound, numPartitions, connectionProperties)
Partition column type should be numeric, date, or timestamp. Check this pull request for details.
Check spark-jdbc-partition.py file for more details.
But what if you don't want to read the whole table's content? 😟
Instead of a full table, you could also use a subquery in parentheses for dbtable option.
For example, (select c1, c2 from table1 where c3 > 40) as s
Or we can use query option. For example,
select c1, c2 from table1 where c3 > 40
Check spark documentation for more details.
spark.read().orc(path)
spark.read().parquet(path)
spark.read().text(path)
There are various modes while saving the dataset
df.write().mode(saveMode)
SaveMode can have these values:
- Append
- Ignore
- ErrorIfExists (default)
- Overwrite
df.write().json(path)
df.write().csv(path)
df.write().jdbc(url, table, properties)
df.write().orc(path)
df.write().parquet(path)
df.write().text(path)
df.write().saveAsTable(tableName);
The goal of Project Tungsten is to improve Spark execution by optimizing Spark jobs for CPU and memory efficiency.
Following optimizations are done be Tungsten:
-
Off-Heap Memory Management using binary in-memory data representation (aka Tungsten row format) and managing memory explicitly.
-
Cache Locality which is about cache-aware computations with cache-aware layout for high cache hit rates.
-
Whole-Stage Code Generation (aka CodeGen).
Project Tungsten uses sun.misc.unsafe API for direct memory access to bypass the JVM in order to avoid garbage collection.
If you persist both dataset and its underlying RDD, you will see a significant difference in the storage size. And the reason is, tugsten makes it smaller in the case of datasets.
Further details can be found here.
Catalyst optimizer is the core component of Spark SQL, which leverages advanced programming language features (e.g. Scala’s pattern matching and quasi quotes) in a novel way to build an extensible query optimizer.
Catalyst contains a general library for representing trees and applying rules to manipulate them. On top of this framework, it has libraries specific to relational query processing (e.g., expressions, logical query plans), and several sets of rules that handle different phases of query execution: analysis, logical optimization, physical planning, and code generation to compile parts of queries to Java bytecode. For the latter, it uses another Scala feature, quasiquotes, that makes it easy to generate code at runtime from composable expressions. Catalyst supports both rule-based and cost-based optimization.
Further details can be found here.
Spark programs can be bottlenecked by any resource in the cluster: CPU, network bandwidth, or memory.
Problems with the Java objects
Java objects are fast to access, but can easily consume a factor of 2-5x more space than the “raw” data inside their fields.
This is due to object header (which is about 16 bytes), UTF-16 encoding for string (stores each string character in 2 bytes), a collection of primitive types are stored as boxed objects
The first way to reduce memory consumption is to avoid the Java features that add overhead, such as pointer-based data structures and wrapper objects. There are several ways to do this:
- Design your data structures to prefer arrays of objects, and primitive types, instead of the standard Java or Scala collection classes (e.g. HashMap). The fastutil library provides convenient collection classes for primitive types that are compatible with the Java standard library.
- Avoid nested structures with a lot of small objects and pointers when possible.
- Consider using numeric IDs or enumeration objects instead of strings for keys.
- If you have less than 32 GB of RAM, set the JVM flag
-XX:+UseCompressedOopsto make pointers be four bytes instead of eight.
Monitoring Garbage Collector
Measuring the Impact of GC by adding -verbose:gc -XX:+PrintGCDetails -XX:+PrintGCTimeStamps to the Java options
GC tuning flags for executors can be specified by setting spark.executor.extraJavaOptions in a job’s configuration. Next time your Spark job is run, you will see messages printed in the worker’s logs each time a garbage collection occurs.
Note these logs will be on your cluster’s worker nodes (in the stdout files in their work directories), not on your driver program.
Try the G1GC garbage collector with -XX:+UseG1GC. It can improve performance in some situations where garbage collection is a bottleneck.
Serialized RDD Storage
If your objects are too large to efficiently store, they can be stored in serialized form. Serialization will make access time high due to on-the-fly deserialization.
Java serialization:
By default, Spark serializes objects using Java’s ObjectOutputStream framework, and can work with any class you create that implements java.io.Serializable. Java serialization is flexible but often quite slow, and leads to large serialized formats for many classes.
Kryo serialization:
Spark can also use the Kryo library (version 4) to serialize objects more quickly. Kryo is significantly faster and more compact than Java serialization (often as much as 10x), but does not support all Serializable types and requires you to register the classes you’ll use in the program in advance for best performance.
You can set it using conf.set("spark.serializer", "org.apache.spark.serializer.KryoSerializer")
If you don’t register your custom classes, Kryo will still work, but it will have to store the full class name with each object, which is wasteful.
Reasons
- RDD doesn't fit in the memory
- Memory utilized by tasks such as reducer task in groupByKey
- Spark’s shuffle operations (sortByKey, groupByKey, reduceByKey, join, etc) build a hash table within each task to perform the grouping, which can often be large
Fix
- RDD can be serialized
- Level of parallelism can be increased to reduce the input set for each task
- Broadcast variables can be used for sharing large data among various workers
Few other relevant configurations
spark.sql.inMemoryColumnarStorage.compressed (default value - true) and spark.sql.inMemoryColumnarStorage.batchSize (default value - 10000)
There are few other properties like spark.sql.files.maxPartitionBytes(default value - 128 mb) spark.sql.shuffle.partitions (default value - 200), etc. are also helpful to fine tune spark jobs.
- Avoid shuffling whenever possible. During shuffling, all shuffle data must be written to disk and then transferred over the network. Each time shuffling will generate a new stage. Repartition, join, cogroup, and any of the By or ByKey transformations can result in shuffles.
- Avoid using
groupbywhenever possible. Try to get the same results usingreduceby,aggregateby, etc. - Use coalesce if you want to decrease the number of the partitions of the RDD instead of repartition. coalesce is useful because avoids a full shuffle, It uses existing partitions to minimize the amount of data that's shuffled.
- If the small RDD is small enough to fit into the memory of each worker we can turn it into a broadcast variable and turn the entire operation into a so-called map side join for the larger RDD.
- Filter input earlier in the program rather than later.
- Don't use count() if you don't want to return the exact number of rows.
- Use the built-in aggregateByKey() operator instead of writing your own aggregations.
- Prefer
TreeReduceandTreeAggregateinstead ofReduceandAggregatewhile dealing with a large amount of data. Check this gitbook for details. - Don't have too big partitions. Your job will fail due to 2 GB limit.
- Use maven shade plugin and relocation tag to avoid jar conflicts.
If data is significantly large, the number of partitions should be atleast equal to the number of cores of the cluster (practically 2-3 times). 128 MB is good size consideration per partition. If the number of partitions is close to but less than 2000, bump to just above 2000.
As a rule of thumb tasks should take at least 100 ms to execute. If your tasks take considerably longer than that keep increasing the level of parallelism, by say 1.5, until performance stops improving.
Let's start with an example. We have 6 node cluster with 16 core and 64 GB memory each.
Few pointers
- 5 cores per executor are good for max HDFS throughput.
- Leave 1 core for Hadoop/Yarn daemon.
- Leave 1 executor for AM(Application Manager).
- Leave 1GB memory of executor node for OS.
- Leave 7-10% memory on executors for heap overhead.
| Parameter | Value |
|---|---|
| Total cores | 16*6 = 96 |
| Total cores after leaving for hadoop/yarn daemon | 96-6 = 90 |
| Number of executors | 90/5 = 18 |
| Number of executors after leaving for AM | 18-1 = 17 |
| Number of executors on each node | 18/6 = 3 |
| Memory per executor | (64-1)/3 = 21GB |
| Memory per executor after counting off-heap overhead | 21 * (1-0.07) ~ 19GB |
-
Properties set directly on the
SparkConftake the highest precedence, then flags passed tospark-submitorspark-shell, then options in thespark-defaults.conffile. -
For detailed spark configuration, check - https://spark.apache.org/docs/latest/configuration.html
-
Higher-Order functions for complex types
- How to use kryo serializer
- Examples for Hash and Range partitioning
- https://spark.apache.org/docs/latest/index.html
- Learning Spark Book by Matei Zaharia, Patrick Wendell, Andy Konwinski, Holden Karau
- https://umbertogriffo.gitbooks.io/apache-spark-best-practices-and-tuning/content/treereduce_and_treeaggregate_demystified.html
- https://jaceklaskowski.gitbooks.io/mastering-apache-spark/
- https://www.slideshare.net/mobile/cloudera/top-5-mistakes-to-avoid-when-writing-apache-spark-applications